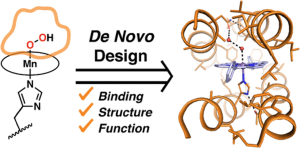
In collaboration with DeGrado (USCF) and Beratan (Duke), the Therien lab looks to uncover the ways that proteins control biological energy transduction by building such function from scratch. Redox proteins transduce energy by orchestrating the movement of electrons, holes, and protons using bound cofactors, redox-active amino acids, titratable sidechains, and buried water. We aim to do much more than to extract electron transfer (ET) parameters from biological systems or to install similar dynamics in small molecules; rather, our aim is to create fully functional de novo proteins that use this recipe to dissect the mechanisms of biological energy transduction in the soft, wet, complex milieu of folded proteins, through both experiment and theory. We will design proteins that independently control the flow of electrons and protons; and we will unify these functions to drive thermodynamically reversible reactions at low overpotential. By binding light-triggerable abiological cofactors, for which we have made numerous recent breakthroughs, our designed proteins will emulate the energy-transduction capabilities of natural proteins and lay groundwork for engineering soft materials that produce exotic photoproducts, possess novel electro-optic function, and transduce energy via innovative pathways.
(1) De novo Design of Crystallographically Characterized Abiological Mn-Porphyrin Binding Protein Capable of Stabilizing High-Valent Mn(V) Species, S. I. Mann, A. Nayak, G. T. Gassner, M. J. Therien, and W. F. DeGrado, J. Am. Chem. Soc. 2021, 142, 252−259. DOI: 10.1021/jacs.0c10136.
(2) Polizzi, N. F.; Wu, Y.; Lemmin, T.; Maxwell, A. M.; Zhang, S.-Q.; Rawson, J.; Beratan, D. N.; Therien, M. J.; DeGrado, W. F., De novo design of a hyperstable non-natural protein–ligand complex with sub-å accuracy. Nat. Chem. 2017, 9, 1157-1164.
Contact Jarrett or Rui for more information on de novo protein design.