Pooled CRISPR sgRNA screening
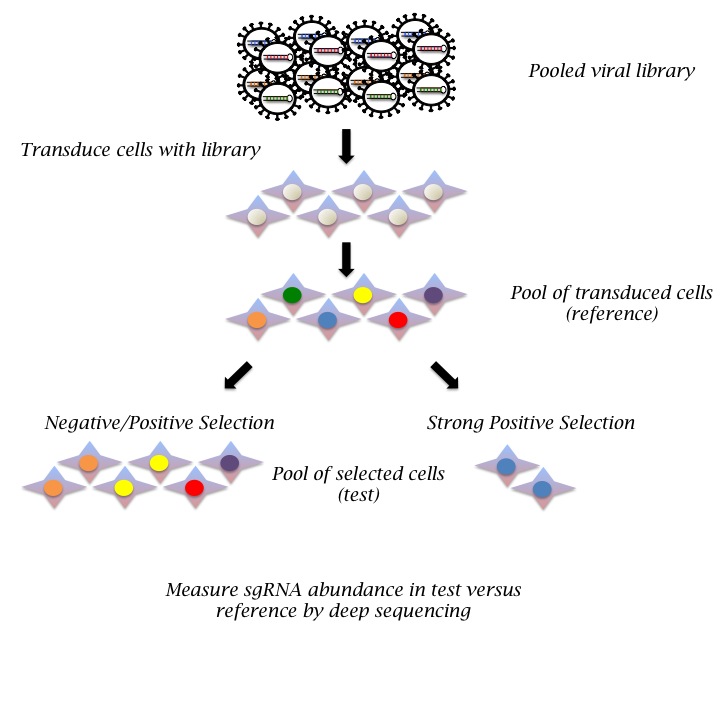
We currently have several genome-scale CRISPR sgRNA libraries available for pooled screening of human or mouse cells. Pooled screens rely on selection strategies that enable separation of cells according to their phenotypes. Effects of the sgRNAs are quantitated as fold enrichment or depletion in the population after selection, see figure below. Screen phenotypes can be based on cell proliferation, such as drug resistance, synthetic lethality or on other selectable features, such as fluorescent reporters or cell size. Of note, pooled screens can utilize complex experimental models that are not amenable to screening in multiwell format, including in vivo animal models.
Libraries:
- Custom libraries available in a variety of vectors
- Human Brunello in lentiCRISPR.v2: 76,441 sgRNAs targeting 19,114 genes
- Human Kinome in lentiCRISPR.v2: 6,104 sgRNAs targeting 763 kinase genes
- Mouse Brie in lentiCRISPR.v2: 78,637 sgRNAs targeting 19,674 genes
- Human CRISPRa (Calabrese): 56,762 (Set A), 56,476 (Set B) lentiviral sgRNA vectors targeting >18,000 transcript isoforms for gain-of-function screening (Sanson et al, Nat. Commun. 2018))
- Human CRISPRi (Dolcetto): 18,901 (Set A), 18,899 (Set B)
For more details, please send inquiries to rnai@duke.edu.
