Comparative Anatomy
Our work is informed by the tenet that structure constrains function, and thus that an understanding of the anatomy of neural circuits can both drive development of hypotheses and support interpretation of data. We choose for each project the experimental methods that will deliver that data we need.
Current projects in the lab make use of:
- tract tracing to visualize connections within and between visual areas, combined with immunohistochemistry to localize modulatory receptors on those connecting axons.
- in situ hybridization to visualize mRNA expression, and immunohistochemistry to visualize protein expression and localization in order to define and identify subtypes of inhibitory neurons.
We collect data using light, epifluorescence, confocal, and electron microscopy and analyze those data using traditional statistics, stereology, and Bayesian approaches.
Current Questions
 In this work, we use single label or multichannel (fluorescence) visualization of mRNA and/or protein expression to quantify populations of cells in the cortex.
In this work, we use single label or multichannel (fluorescence) visualization of mRNA and/or protein expression to quantify populations of cells in the cortex.
We define these populations using a number of different classification schemes, including signaling molecules (e.g. GABA, dopamine, NPY, somatostatin) and calcium-binding protein expression (e.g. parvalbumin, calbindin).
The questions we're interested in answering through this work include:
-
- How similar are cortical circuits? Is there a canonical circuit that holds true across species and brain areas, or are different areas (such as the primary visual cortex, or the lateral intraparietal area) comprised of different relative mixtures of the same neurons, or even entirely different types of neurons?
- What cell types can be identified in cortex? Do these types differ between species?
- What signaling molecules tend to be expressed together within single neurons? How does this expression confer or constrain function?
 Extracellular levels of any given neuromodulator (for example serotonin) are determined by a number of factors, some of which can be assayed from structural studies.
Extracellular levels of any given neuromodulator (for example serotonin) are determined by a number of factors, some of which can be assayed from structural studies.
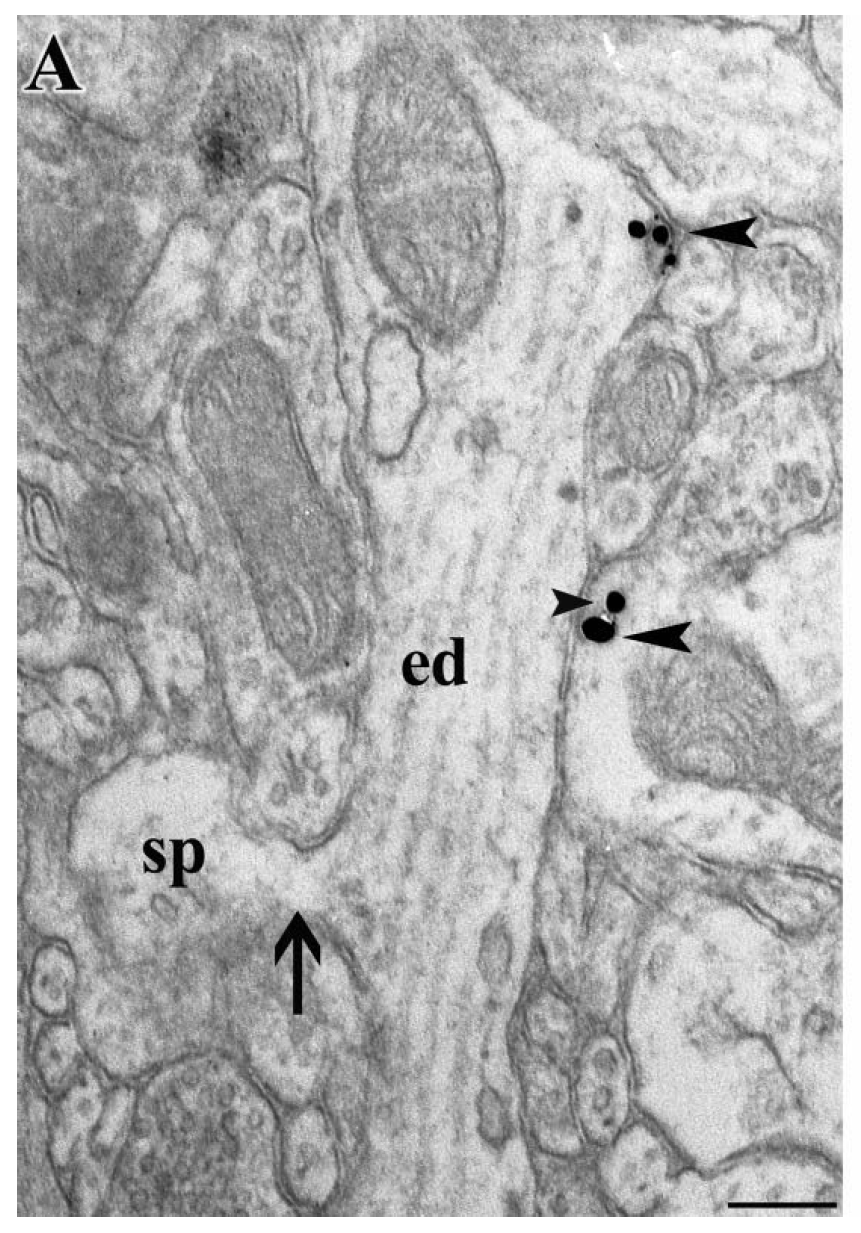
Structural determinants of extracellular concentration will include the density of modulatory axons and the barriers to diffusion through the tissue. We quantify these structural features at the light and electron microscopic features.
Some of the questions we are interested in addressing through this work include:
-
- Over what spatial scales do these structural determinants differ? For example, are there laminar differences in the density of axons?
- Are axonal release sites colocalized with signal termination mechanisms (e.g. transporters that remove modulators from the extracellular space)? What does this tell us about the diffusion?
- What are the structural markers of a release site?
 All brain circuits are subject to neuromodulation, but it is often the case that only some of the cells and cellular compartments in that circuit are actually 'listening' to those modulatory signals.
All brain circuits are subject to neuromodulation, but it is often the case that only some of the cells and cellular compartments in that circuit are actually 'listening' to those modulatory signals.
We can identify the signal receivers for a circuit by probing for the expression of receptors, and this information both constrains hypotheses about functional effects of modulator release, and help us interpret the effects we see in our in vivo experiments.
Some of the questions we are asking in this space are:
-
- Which pathways into and through a given cortical area (i.e. feedfoward, feedback, local circuit) are targets for neuromodulation?
- Are there species differences in the expression of modulatory receptors?
- Are there differences within a species in the ways receptors are expressed across cortical areas and layers?
- On what subcellular compartment(s) are modulatory receptors localized?




