A Virtual Symposium hosted by the Fitzpatrick Institute for Photonics | May 16, 2020
Special Topics: Photonics in the Era of Data Science: From Smart Sensing to AI
Virtual Poster Session
Click a poster thumbnail image to enlarge.
Awardees of 2020 FIP Virtual Poster Session
1st Place – Kristen Hagan
Poster#21 “Wavefront sensorless handheld multimodal
adaptive optics scanning laser ophthalmoscope”
2nd Place – Natalie Rozman
Poster#8 “High-Quality Factor THz Metamaterial”
3rd Place – Yang Liu
Poster #12 “Synergistic Immuno Photothermal Nanotherapy (SYMPHONY)
for Effective Cancer Treatment”
Honorable Mention – Shangguo Hou
Poster#6 “Real-time 3D Molecule Tracking Microscopy”
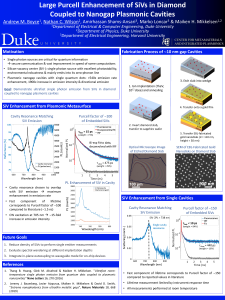
1. Large Purcell Enhancement of SiVs in Diamond Coupled to Nanogap Plasmonic Cavities
View abstract
Single photon sources could be crucial components of quantum computing and quantum information systems. Naturally occurring single photon sources are insufficient for many applications because of their low photon count rates, short coherence times and lack of a directional emission pattern. A novel nanostructure, the nanopatch antenna (NPA) consists of metallic nanoparticles separated from a metallic film by a nanoscale gap layer and has demonstrated the largest emission rate enhancements of spontaneous emitters via the Purcell effect to date, including 540-fold enhancement of single photon emission from colloidal quantum dots. However, applications of colloidal quantum dots are limited due to their susceptibility to photobleaching and degradation over time. On the other hand, color centers in diamond offer long coherence times and excellent photostability. Thus, combining these color centers with NPAs could yield robust single photon sources with high photon count rates and directional emission. Our approach for integrating diamond color centers into the NPA involves etching single-crystal diamond slabs with implanted silicon vacancies (SiVs). Nanoparticles are fabricated by EBL and transferred onto the etched diamond slab on gold using a PDMS stamp. Fluorescence lifetime measurements were carried out by using pulsed laser excitation at 514 nm. SiV emission was detected by low-jitter avalanche photodiodes and tagged using a high-resolution time to digital converter. Fast decay components were measured on many different spots on the sample that indicated instrument response limited lifetimes on the order of 10 ps, corresponding to ~100-fold enhancement compared to the intrinsic lifetime of SiVs.
2. Programming temporal DNA barcodes for single-molecule fingerprinting
Shalin Shah 1,2, Abhishek Dubey 2,3 and John Reif 1,2
1 Department of Electrical and Computer Engineering, Duke University, Durham, NC, United States – 27701
2 Department of Computer Science, Duke University, Durham, NC, United States – 27701
3 Computational Sciences and Engineering Division, Health Data Sciences Institute, Oak Ridge National Lab, Oak Ridge, Tennessee 37831, United States
View abstract
Fluorescence microscopy offers a versatile solution to study the dynamics of biology at the mesoscale. An important challenge in the field is the simultaneous study of several objects of interest, referred to as optical multiplexing. For improved multiplexing, some prior techniques used repeated reporter washing or the geometry of nanostructures; however, these techniques require complex nanostructure assembly, multiple reporters, or advanced multistep drift correction. Here we propose a time-based approach, for improved optical multiplexing, that uses readily available inexpensive reporters and requires minimal preparation efforts. We program short DNA strands, referred hereby as DNA devices, such that they undergo unique conformation changes in the presence of the dye-labeled reporters. The universal fluorescent reporter transiently binds with the devices to report their activity. Since each device is programmed to exhibit different hybridization kinetics, their fluorescent time trace, referred to as the temporal barcode, will be unique [1]. We model our devices using continuous-time Markov chains and use stochastic simulation algorithm to generate their temporal patterns [2]. We first ran simulation experiments with a small number of DNA devices, demonstrating several distinct temporal barcodes, all of which use a single dye color. Later, using nanostructure-based devices, we designed a much larger pool of temporal barcodes and used machine learning for classification of these barcodes. Using our simulation experiments and design principles we also experimentally demonstrate our DNA devices. We show DNA strands can be programmed for generating a multitude of uniquely identifiable molecular barcodes. Our technique can be easily incorporated with the existing orthogonal methods that use wavelength or geometry to generate a large pool of distinguishable molecular barcodes thereby enhancing the overall multiplexing capabilities of single-molecule imaging [3].
3. A Functional in vitro model of human kidney glomerulus
Xingrui Mou1, Rohan Bhattacharya1, Bowen Sun2, Po-Chun Hsu3, Samira Musah1,4,*
1Department of Biomedical Engineering,
2Department of Materials Science and Engineering,
3Department of Mechanical Engineering and Materials Science,
4Department of Medicine, Division of Nephrology, Duke University, Durham, NC, USA.
*Corresponding author: email, samira.musah@duke.edu
View abstract
The kidney is an essential organ for the maintenance of health and homeostasis through filtration of blood and production of hormones to regulate organ function. Kidney disease has rapidly become a public health crisis as a growing number of patients develop the disease with subsequent progression to end stage kidney disease and organ failure. Efforts to study the etiology and progression of kidney diseases include the application of two-dimensional (2D) cell culture plates and animal models. However, challenges remain as 2D cell culture systems lack the three-dimensional (3D) tissue structure and organization that play key roles in mediating cellular crosstalk and cell-matrix interactions. Furthermore, animal models cannot accurately recapitulate human physiology due to species-specific differences and divergent developmental and cell signaling pathways. Therefore, we aim to engineer a capillary-like 3D in vitro system that mimics the human kidney glomerulus by applying electrospinning and stem cell differentiation technologies. The human kidney glomerulus is a capillary-rich structure that serves as the primary site for blood filtration and it is the target of various forms of kidney diseases. Specifically, we are developing 3D matrices coupled with microfluidic systems to mimic the structure and blood filtration functions of the human kidney glomerular capillaries. We will integrate these engineered scaffolds with human glomerular podocytes obtained through our previously reported stem cell differentiation method and an endothelial cell differentiation protocol to generate kidney cells. The resulting in vitro system could serve as a platform for mechanistic studies and therapeutic discoveries for human kidney diseases.
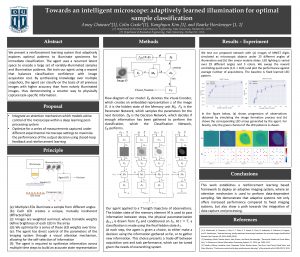
4. Towards an intelligent microscope: adaptively learned illumination for optimal sample classification
View abstract
Recent machine learning techniques have dramatically changed how we process digital images. However, the way in which we capture images is still largely driven by human intuition and experience. This restriction is in part due to the many available degrees of freedom that alter the image acquisition process (lens focus, exposure, filtering, etc.). Here we focus on one such degree of freedom – illumination within a microscope – which can drastically alter information captured by the image sensor. We present a reinforcement learning system that adaptively explores optimal patterns to illuminate specimens for immediate classification. The agent uses a recurrent latent space to encode a large set of variably illuminated samples and illumination patterns. We train our agent using a reward that balances classification confidence with image acquisition cost. By synthesizing knowledge over multiple snapshots, the agent can classify on the basis of all previous images with higher accuracy than from naively illuminated images, thus demonstrating a smarter way to physically capture task-specific information.
5. The Effect of Magnetic Fields on Plasmon Resonance Frequencies of Metal Nanostructures
View abstract
Magnetic fields are demonstrated to affect surface plasmon-polaritons on non-magnetic Au nanostructures. Previously, the magnetooptical Kerr and Faraday polarization dependent effects have been observed on hybrid magnetic-plasmonic structures, and it has been theoretically demonstrated that under high magnetic field strength (>10T) a splitting of the plasmon absorption band occurs with circularly polarized light. Here, the surface plasmon-polariton absorption by Au nanostructures are measured utilizing a magnetospectrometer with a magnetic field force up to 1.5T and light with varied polarization. It is found that as the magnetic field strength increases, the energy and the lifetime of the plasmon resonance are altered. Coupled with theoretical work, our findings suggest a Lorentz force is acting directly on the elections under a magnetic field causing changes to surface plasmon polariton. Though providing further insight into surface plasmons, this work grants access to increased understanding across the fields of plasmonic catalysis, sensing, and optoelectronics.
6. Real-time 3D Single Molecule Tracking Microscopy
View abstract
Single molecule tracking has the potential to revolutionize the study of biological systems. However, most current single molecule imaging and tracking methods are limited to two dimensions. Meanwhile, real-time 3D single molecule tracking methods, which use active feedback to “lock-on” to single molecules in solution have until now been limited to observation times of 500 ms or less, greatly limiting their utility. Recently we developed a real-time 3D single particle tracking method known as 3D Dynamic Photon Localization Tracking (3D-DyPLoT). By implementing optimized tracking parameters based on single molecule model simulation, in here we present a 3D single-molecule active real-time tracking method (3D-SMART) which is capable of track single fluorophores in solution for several minutes at a time, two orders of magnitudes longer than previously reported methods, with photon limited temporal resolution. We demonstrate the application of 3D-SMART to track the free, 3D diffusion of DNA, RNA and proteins in solution. With two-color observation channels, 3D-SMART was also applied to observing the transcription of single DNA molecules in solution by monitoring the real-time production of mRNA. This real-time 3D single molecule tracking method promises to be powerful tool for capturing the dynamics of single biomolecules at high speeds and over 3D distances.
7. Designing a task-specific microscope using a deep neural network to improve image classification accuracies
View abstract
Traditionally, microscopes are optimized to create high-resolution images over a large area for visual inspection, which often leads to bulky optics and expensive sensors. In many scenarios, machine learning (ML) algorithms do not necessarily need a well-resolved image for their inference tasks – instead, an optimally coded low-resolution image can contain a sufficient amount of information for high ML decision accuracy. Here, we present a framework to design a microscope, using a novel deep neural network, which can jointly optimize the microscope’s hardware for improved ML performance. In this work, we use supervised learning to optimize both the pupil transmission function and the pattern of sample illumination, which is otherwise a challenging optimization problem due to the large number of constraints involved. The pupil transmission and illumination patterns are sample-specific and can be easily updated to optimally classify alternative sample categories in a simple, low-cost, 4F microscope, by changing the pupil plane mask and the illumination pattern on a programmable LED array. Our preliminary results show that by jointly designing imaging system hardware with a deep network software, we can develop low-resolution wide-field microscopes that can provide higher task-specific classification throughput and thus faster diagnoses than alternative approaches.
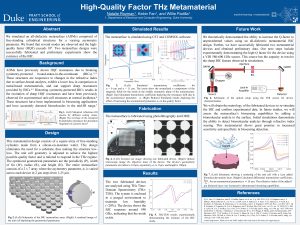
8. High-Quality Factor THz Metamaterial
View abstract
The overarching objective of the proposed research is to investigate the feasibility of an all-dielectric metasurface with high specificity and sensitivity for biosensing applications. Metamaterials can be tailored to have an extremely precise response in a desired frequency range. Using this innovative structure, I will create a photonic biosensor for Ebola virus disease detection. Through understanding ofthe physics underlying the existence of Bound-States-in-the-Continuum (BICs), I have performed initial simulations verifying designs which may be used to carry out the proposed research. An ideal structure consists of all-dielectric metasurfaces (ADMs) comprised of a free-standing square array of silicon cylinders attached with thin fins for a varying asymmetry parameter (α). The geometry of the resonator was altered by removing part of the cylinder by α, while maintaining the periodicity P, and interconnect width w, as depicted in the inset in Fig. 1. This asymmetry gives rise to several modes that result in an unprecedented high-Q exceeding 106, observed in the simulated frequency dependent transmission (T(ω)) at f/f_0). Initial simulations were carried out, verifying that this structure can be designed for the near-infrared range for photonic biosensing applications. Thus the high spectral resolution of this ADM structure shows great promise for metamaterial applications in biosensing detection.
9. Passive Cavitation Mapping in Shockwave Lithotripsy
View abstract
Shockwave lithotripsy enables non-invasive therapy for kidney stone patients. Cavitation generated by the tensile shockwaves plays an important role in stone erosion. However, it is also the major reason that causes tissue damage and renal hemorrhage due to the violent collapses of cavitation bubbles. It is therefore crucial to acquire temporal and spatial distributions of cavitation collapsing to predict possible hemorrhage occurrences during the shockwave treatment. Conventional cavitation detection and mapping approaches, including photography, active cavitation mapping (ACM) and passive cavitation mapping (PCM), provide incomplete cavitation collapse information of individual bubbles for biological tissue and degrade the precision of localizing hemorrhages during the treatment due to the temporal randomness of cavitation collapsing. To image the bubble collapse distributions more precisely, a sliding-window passive cavitation mapping algorithm is proposed and validated. By sliding the reconstruction window and searching along time, correct time origin for each individual bubble collapses can be identified. Laser-induced single bubbles were generated in free water space and in a transparent plastic tube, respectively, and the cavitation collapse signals were captured and reconstructed using the proposed method. High-speed camera imaging and active cavitation mapping were performed simultaneously as ground-truth to validate the accuracy of passive method. Furthermore, lithotripter shockwave induced cavitation were acquired using the same methods. Images from all three modalities match well and the overall experiments demonstrated the reliability and accuracy of the proposed cavitation mapping method.
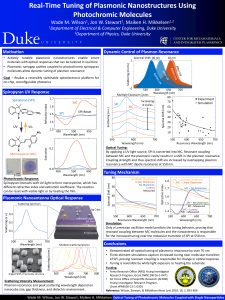
10. Real-Time Tuning of Plasmonice Nanostructures Using Photochromic Molecules
Wade M. Wilson1,2,†, Jon W. Stewart1,2,† and Maiken H. Mikkelsen1,2,3,*
1Center for Metamaterials and Integrated Plasmonics, Duke University, Durham, NC 27708
2Department of Electrical and Computer Engineering, Duke University, Durham, NC 27708
3Department of Physics, Duke University, Durham, NC 27708
*Correspondence to: m.mikkelsen@duke.edu, +1 (919) 660-0185
†These authors contributed equally to this work.
View abstract
Plasmonic nanostructures with actively tunable optical properties could realize beam steering surfaces, on-chip hyperspectral detectors and dynamically reconfigurable optoelectronic devices. A variety of tunable materials have been integrated with plasmonic structures, however, the tuning range in the visible regime has been limited to less than the linewidth of the resonance, resulting in small on/off ratios critical for these applications. Here we demonstrate all optical, real-time tuning of plasmon resonances up to 71 nm in the visible regime by incorporating spiropyran into plasmonic nanopatch antennas. The tuning mechanism is elucidated by comparing experimental data to finite-element simulations using different models for the dispersion in the photochromic film. This comparison reveals that the spectral shift is due to resonant coupling between spiropyran and the plasmonic nanoantennas.
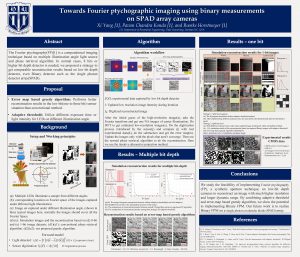
11. Towards Fourier ptychographic, imaging using binary measurements on SPAD array cameras
View abstract
Recently developed Single-photon Avalanche Diode (SPAD) array cameras have single photon sensitivity and can provide time-of-flight information for LIDAR imaging. These SPAD cameras, however, have very few pixels and readout binary images, which are typically averaged to provide an image with sufficient dynamic range. Here, we propose to implement a modified version of Fourier ptychography (FP), a synthetic aperture technique, on SPAD cameras to reconstruct an image with much higher resolution and larger dynamic range from its binary measurements. We successfully validate this using simulated and experimental results to show its potential for recording LIDAR images at high resolution and speed.
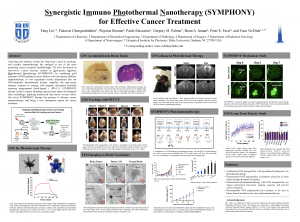
12. Synergistic Immuno Photothermal Nanotherapy (SYMPHONY) for Effective Cancer Treatment
Yang Liu1, Pakawat Chongsathidkiet2, Paolo Maccarini1, Gregory M. Palmer3, Peter E. Fecci4, and Tuan Vo-Dinh1,5,6*
1. Duke University Biomedical Engineering Department
2. Duke University Medical Center Pathology Department
3. Duke University Medical Center Radiation Oncology Department
4. Duke University Medical Center Neurosurgery Department
5. Duke University Chemistry Department
6. Duke University Fitzpatrick Institute for Photonics
* Corresponding author: tuan.vodinh@duke.edu
View abstract
Glioblastoma (GBM) is the most common and aggressive primary brain cancer and it has more than 10,000 newly diagnosed patients in the United States each year. Even with the highest first-year cost (> $120,000), the prognosis for GBM patients is dismal and the median survival is only 15 months after aggressive treatments including surgery, chemotherapy, radiation therapy, and targeted drug therapy. Less than 5% of the patients survive for more than 3 years. Despite enormous efforts, there have been no major treatment advances in over two decades, and GBM is still a devastating disease with essentially 100% mortality. Therefore, there is a clear and urgent clinical need to develop novel and effective therapeutic methods to dramatically improve GBM treatment efficacy. We have developed an innovative cancer therapy named as Synergistic Immuno Photothermal Nanotherapy (SYMPHONY) by combining gold nanostars (GNS)-mediated photothermal ablation with checkpoint inhibitor immunotherapy. GNS nanoparticle has multiple sharp branches for tip-enhanced plasmonics and tunable plasmonic absorption in the near-infrared (NIR) tissue optical window. GNS has superior photonto- heat conversion capability for effective photothermal therapy with NIR laser. Our group has innovated a toxic chemical-free method to synthesize GNS nanoparticles and apply the biocompatible GNS nanoparticles for SYMPHONY therapy. In vivo experiment with brain cancer murine animal models demonstrate that our superior GNS-mediated photothermal therapy dramatically amplifies the anticancer immune response in synergy with checkpoint blockade immunotherapy. SYMPHONY therapy results in not only primary tumor shrinkage but also recurrence prevention, implying the generation of an anti-cancer vaccine effect. As a result, our novel SYMPHONY therapy has the potential to substantially improve outcomes of brain cancer patients in future clinical applications.
13. Differential phase contrast imaging on a gigapixel microscope
View abstract
Asymmetric illumination-based differential phase contrast (AIDPC) is a full-field phase-gradient imaging method that can capture phasegradient information at microscopic resolution. However, currently explored differential phase contrast systems image over a small field of view. In this work, we apply AIDPC to images acquired from an array of 96 imagers that exhibit partially overlapping fields-of-view, enabling high-resolution (10 um) phase gradient measurements over hundreds of square centimeters. We present experimental results showing how our multi-camera AIDPC approach recovers high-quality surface normals from both diffuse and specular macroscopic objects.
14. Particle-by-particle in situ characterization of the protein corona via real-time 3D single particle tracking microscopy
View abstract
Engineered nanoparticles adsorb proteins when they are exposed to biological fluids, forming a dynamic protein corona that consists of tightly bound “hard” corona, and rapidly exchanged “soft” corona. This full protein corona alters the surface identity of nanoparticles, affecting their behavior in biological systems. However, a thorough understanding of the protein corona is lacking due to the insufficiency of current ensemble techniques. Here, we introduce real-time single 3D particle tracking microscopy to “lock-on” to single freely diffusing polystyrene nanoparticles (PSNPs) and probe their individual protein coronas in solution. Using this method, we study the protein corona content of individual PSNPs in solutions containing bovine serum albumin (BSA), immunoglobulins (Ig) and fetal bovine serum (FBS). Using mean squared displacement measurements combined with real-time protein fluorescence, we quantified “hard” corona growth particle-by-particle. Furthermore, we applied a lock-in type frequency filtering method to extract the full protein corona signal (i.e. both “soft” and “hard” corona) in high background environments. Our method enables the study of individual protein coronas without removing the particle or the protein from its native solution, opening up the possibility to study transient and dynamic protein-nanoparticle interactions which are destroyed by current bulk techniques.
15. Advanced Light Imaging and Spectroscopy (ALIS) at Duke
View abstract
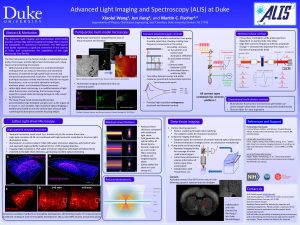
The Advanced Light Imaging and Spectroscopy (ALIS) facility provides cutting-edge optical imaging technology that is beyond the capabilities of commercial instruments. The first instruments in the facility include: a lattice light-sheet microscope and a deep-tissue serial sectioning microscope. The lattice light sheet microscope allows us to image 3D dynamics for hundreds of volumes at the diffraction limit and beyond with super low photoxicity. The strength of this technology lies in the combination of high spatial resolution and high acquisition speed. This microscope is ideally suited to image a range of biological samples, ranging from intracellular components to small organisms (such as fruit fly embryos). The deep-tissue serial sectioning microscope operates at the other end of the spatial scale: large biological samples (such as organs of a mouse). In such samples, high-resolution optical imaging is traditionally limited to the surface layer because light scattering and absorption prevents focusing deeper. Our approach is to repeatedly image a small (few hundred μm thick) layer of the sample at a time and then remove the imaged section to expose deeper layers. Fully automated imaging and sectioning allows for gap-less, high-resolution imaging of large volumes. The 980-square-foot facility represents a significant investment in the sciences at Duke and complements the capabilities of the Light Microscopy Core Facility, it will advance research of many biological and medicine groups significantly.
16. Chalcogenide-based Photonic Quasicrystals for Novel Phase Matching
Jiannan Gao1, Wiktor Walasik1, Mikhail Shalaev1, Jesse Frantz2, Jason D. Myers2, Robel Y. Bekele3, Jasbinder S. Sanghera2, and Natalia M. Litchinitser1
1Department of Electrical and Computer Engineering, Duke University, Durham, North Carolina, 27708, USA
2US Naval Research Laboratory, 4555 Overlook Ave., SW, Washington, DC 20375
3 University Research Foundation, Greenbelt, MD, USA 20770, USA
View abstract
Quasicrystal-based engineered photonic nanostructures are unique structures with long-range order but no periodicity. Quasicrystals are a class of lattices characterized by a lack of translational symmetry but obeying rotational symmetry. Similar to naturally occurring quasicrystals, due to their unique symmetry properties, artificial quasicrystals are expected to possess properties that are different from both crystals and glasses. In this work, we focus on novel regimes of nonlinear light-matter interactions enabled by engineered photonic quasicrystals. In particular, in the field of nonlinear optics, the advantage of using PQCs is the possibility of supporting unlimited combinations of wavevectors that may appear in their reciprocal lattices. We report (i) the nonlinear interactions engineering using two approaches – dispersion engineering utilizing the strongly modified dispersion relations in the vicinity of the bandgaps of the PQC and quasi-periodic quasi-phase matching; (ii) design and fabrication of PQCs in chalcogenide waveguides; and (iii) experimental demonstration of the predicted new phase-matching regimes enabled by the PQCs. These structures are realized in arsenic trisulfide (As2S3) chalcogenide glass that displays a very good nonlinear figure of merit in both the near-infrared and the mid-wave infrared spectral bands. Here, we show that the proposed structures realized in such a highly nonlinear medium may enable a promising platform for both fundamental studies of nonlinear light-matter interactions and applications in wavelength conversion, supercontinuum generation, and development of classical and quantum optical sources.
17. Quantitative Phase Imaging provides mechanical phenotyping which aligns with Atomic Force Microscopy
View abstract
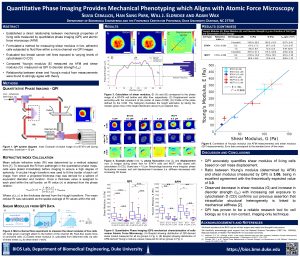
Current standard methods for probing cellular stiffness are slow, difficult, and utilize complex or indirect detection schemes which limit their ability to conduct large scale, or longitudinal studies of cellular mechanics. To enable direct, non-invasive measurement of cellular mechanical properties, we previously developed a quantitative phase imaging (QPI) based method to measure cellular deformation in response to shear stress. The primary mechanical parameters we used to discriminate between different cell and cancer types was shear stiffness, analogous to a spring constant. Using this approach, we studied changes in cellular mechanical integrity due to pharmacological treatment and heavy-metal toxicity. Now, we have established a direct relationship between QPI-derived mechanical parameters and atomic force microscopy (AFM), the gold standard for measuring cellular viscoelastic characteristics. We formulate a method for measuring shear modulus in live, adherent cells from QPI images of cells subjected to shear flow within a flow cell. To validate the approach, we evaluate two breast cancer cell lines exposed to various levels of cytochalasin D, an actin depolymerizing toxin. Each group is characterized by a commercial AFM to measure Young’s modulus and analyzed using our QPI system to quantify shear modulus. We also compare both of these mechanical parameter to disorder strength, a phase fluctuation metric that reveals structural homogeneity. Relationships between shear modulus and Young’s modulus measurements are found to strongly agree with theory, confirming the validity of previous mechanical characterizations using QPI, indicating the utility of QPI as a powerful tool for cellular mechanical studies.
18. A Multi-Resolution Approach to Environment Visualization for Real-Time Active Feedback Single Particle Tracking
View abstract
Real-Time active feedback single particle tracking using the 3D-DyPLoT method is a powerful tool for locking-on to moving nanoscale particles and obtaining quantitative information about their dynamics. However, the tracking system alone is unable to determine the full nature of interactions between a particle and its environment due to the limited field of view required to track particles moving at high speeds. Such interactions, as between a virus infecting a cell, underly many interesting biological questions. The integration of a 3D-FASTR two-photon laser-scanning microscope into the pathway of the 3D-DyPLoT system enables the acquisition of 3D volumes acquired simultaneously during particle tracking. This capability results in the visualization of co-registered 3D volumes with trajectories that show the particle’s motion relative to the volume over time.
19. High-sensitivity pump-probe microscopy based on MHz time-delay modulation
View abstract
We present a new imaging method for femtosecond pump-probe microscopy that explores time-delay modulation at a MHz rate. This method not only greatly reduces the background signal but also enables the real-time calculation of differences computation of images with respect to the inter-pulse delay. We experimentally demonstrate this method by imaging perovskite solar cell materials as well as fruit fly larvae, and demonstrate the real-time differencing on melanin nanoparticles and mice skin samples.
20. Crossed-beam pump-probe microscopy
View abstract
We present a new imaging method for pump-probe microscopy that explores non-collinear excitation. This method (crossed-beam pump-probe microscopy, or CBPM) can significantly improve the axial resolution when imaging through low-NA lenses, providing an alternative way for depth resolved, large field-of-view imaging. We performed a proof-of-concept demonstration, characterized CBPM’s resolution using different imaging lenses, and measured an enhanced axial resolution for certain types of low-NA lenses.
21. Wavefront sensorless handheld multimodal adaptive optics scanning laser ophthalmoscope
Kristen Hagan1, Theodore B. DuBose1, David Cunefare1, Corey Simmerer1, Gar Waterman1, Jongwan Park1, Anthony N. Kuo1,2, Ryan P. McNabb2, Joseph A. Izatt1,2, and Sina Farsiu1,2
1Department of Biomedical Engineering, Duke University, Durham, North Carolina;
2Department of Ophthalmology, Duke University School of Medicine, Durham, North Carolina
View abstract
Adaptive optics scanning laser ophthalmoscopy (AOSLO) has enabled in vivo imaging of individual cone and rod photoreceptors in the human eye, revolutionizing the study of human retinal structure, function, and disease states by vision scientists and physicians. AOSLOs are usually implemented as tabletop systems, thereby preventing patients who cannot sit upright and/or fixate for extended time periods from being imaged by these high-resolution systems. Previously, we addressed this limitation and extended AOSLO to excluded patient populations by designing and fabricating the first confocal handheld AOSLO (HAOSLO) capable of cone photoreceptor visualization in both supine adults and neonates. However, recent studies have shown that confocal AOSLO images contain imaging artifacts and cannot obtain information about comprised cone structure in diseased eyes, leading to ambiguous or misleading patient data. These studies have demonstrated that the collection of multiply backscattered light into two offset non-confocal split-detection (SD) channels can allow for a more reliable view of the patient retina by imaging the cone inner segment. In this project, we introduce the extension of our HAOSLO probe for SD and confocal imaging of cone photoreceptors in the first ever multimodal (M)-HAOSLO probe. Confocal and SD images revealing cone photoreceptors were collected in a tabletop implementation as well as in M-HAOSLO’s handheld operation mode from healthy adult subjects. Besides the unprecedented miniature form factor and low weight, this is the first portable SD AOSLO system as well as the first system to collect SD cone inner segment images following sensorless optimization of the wavefront.
22. Mapping Solvation Heterogeneity in Live Cells by Hyperspectral Stimulated Raman Scattering Microscopy
View abstract
Water has long been known not just to be a spectator, but an active participant in biochemical processes. The local structure and dynamics of intracellular water constitute the cornerstone for understanding cellular function. Fundamentally, direct visualization of subcellular solvation heterogeneity is essential but remains challenging with commonly used NMR methods due to poor spatial resolution. The classical physical chemistry tool to probing solvation in molecules is the use of vibrational frequency as a readout of the molecular environment, particularly using vibration sensors such as nitriles. However, this approach has not been successfully applied to map intracellular solvation due to the combination of small peak shifts and small Raman scattering cross-sections. Here, we boost the Raman scattering signal using a near-resonant nitrile-containing molecule (Rhodamine 800) with hyperspectral stimulated Raman scattering (hsSRS). The near-resonant condition of Rhod800 enables a 200-fold enhancement of the effective Raman scattering crosssection, allowing solvation sensing in the solution phase, micron-scale droplets, and cellular environments. Through calibration, we quantify the degree of solvation between the cytoplasm (29.5%, S.E. 1.8%) and the nucleus (57.3%, S.E. 1.0%) and further validate this heterogeneity is related to the macromolecular crowding within the cytoplasm. These studies represent a new picture of the water structure within cells, wherein there is less “bulk” or “free” water available for participating in hydrogen-bonding and solvation. This work opens up new avenues to explore environmental variance in complex systems with high spatiotemporal resolution.
23. Applications of Nonlinear Optical Spectra and Imaging in Cultural Artifacts Science
View abstract
Pump-probe microscopy is a powerful technique to improve the evaluation of artworks. Conventional research on paintings usually relied on the physical-removed samples from the artworks, which cannot provide depth information. In pump-probe microscopy, with the non-linear optical interactions between the ultrafast, pulsed laser beams and paints, high-resolution, three-dimensional images of paint layers can be obtained without physical removal of samples. By the help of this technology, subsurface paint-layer analysis can be achieved non-invasively with structural and molecular contrast in a 14th century painting; in addition, the light-induced degradation of vermilion, a red historical pigment, was investigated and the degradation products were successfully differentiated and mapped on micro-scale. In the future, the degradation of cadmium sulfide (CdS), a popular yellow pigment, will be imaged under pump-probe microscopy and the potential degradation mechanisms will be studied. Here we provide an update on our spectroscopic and microscopic techniques and describe applications to the study of cultural art objects.
24. SERS-based Plasmonic Coupling Interference (PCI) Nanoprobes for Multiplex Detection of MicroRNA Cancer Biomarkers
Hsin-Neng Wang1,2, Bridget M. Crawford1,2, Stephen J. Norton1,2, Tuan Vo-Dinh1,2,3(*)
1. Departments of Biomedical Engineering, Duke University, Durham, NC 27708, USA
2. Fitzpatrick Institute for Photonics, Duke University, Durham, NC 27708, USA
3. Department of Chemistry, Duke University, Durham, NC 27708, USA
View abstract
MicroRNAs (miRNAs), small non-coding endogenous RNA molecules, are emerging as promising biomarkers for early detection of various diseases and cancers. Practical screening tools and strategies to detect these small molecules are urgently needed in order to facilitate the translation of miRNA biomarkers into clinical practice. In this work, a label-free biosensing technique based on surfaceenhanced Raman scattering (SERS), referred to as “plasmonic coupling interference (PCI)”, was applied for the multiplex detection of miRNA biomarkers. The sensing mechanism of the PCI technique relies on the formation of a nanonetwork consisting of nanoparticles with Raman labels located between adjacent nanoparticles that are interconnected by DNA duplexes. Due to the plasmonic coupling effect of adjacent nanoparticles in the nanonetwork, the Raman labels exhibit intense SERS signals. Such effect can be modulated by the addition of miRNA targets of interest that act as inhibitors to interfere with the formation of this nanonetwork, resulting in a diminished SERS signal. In this study, the PCI technique is theoretically analyzed and the multiplex capability for detection of multiple miRNA cancer biomarkers is demonstrated, establishing the great potential of PCI nanoprobes as a useful diagnostic tool for medical applications.
25. Probing the Spatial Heterogeneity of Photoexcitation Dynamics in Perovskite Thin Films with Femtosecond Time-Resolved Nonlinear Optical Microscopy
View abstract
Hybrid organic-inorganic metal halide perovskites (PVSKs), known as the next generation solar cell materials, has been developed rapidly in the past five years. Its power conversion efficiency (PCE) rises from 3.6% to more than 22.1% and its stability is also improved. Although many studies of its fabrication methods and PCE are reported, its fundamental photoexcitation dynamics is still unclear. To investigate its photoexcitation dynamics, conventional time-resolved analysis tools, such as time-resolved transient absorption (TA) spectroscopy and time-resolved microwave conductivity, are often used. But they can only provide little spatial information of PVSK. Here, we demonstrate pump-probe microscopy is able to reveal photoexcitation dynamics in PVSKs on the femto- to picosecond timescale and also provide sub-micro spatial resolution. We investigate crystalline CH3NH3PbI3 perovskite thin layers on glass substrates and sort the photoexcitation dynamics by different nonlinear processes it causes, such as two photon absorption, ground state depletion and excited state absorption. We combine time-resolved photoluminescence (TRPL) images and pump-probe images to visualize the spatial heterogeneity of photoexcitation dynamics.
26. Photoacoustic Imaging of tattoo inks and evaluation of tattoo treatment
View abstract
Photoacoustic imaging (PAI), as a new imaging method, has excellent optical contrast and ultrasonic spatial resolution. PAI is being extensively studied in clinical settings for biomedical diagnostic applications in vivo. In this poster, we illustrate the utility of PAI as a non-invasive tattoo imaging tool and evaluate the effectiveness of laser tattoo removal therapy. We performed an extracorporeal photoacoustic performance test on 6 different tattoo ink samples embedded in porcine skin before and after laser treatment, and we quantitatively determined the effect of laser treatment. By comparing and quantifying tattoo skin samples before and after treatment, we evaluated the effect of laser treatment on tattoo ink. PAI provides excellent contrast, describing the shape and area of the ink on the skin. Our results reveal the photoacoustic properties of tattoo inks and demonstrate the potential clinical value of PAI in intradermal tattoo imaging. PAI can be used as a preoperative clinical aid in assessing tattoos and may guide the process of laser removal treatment.
27. Fiberoptics SERS Sensors using Plasmonic Nanostar Probes for Detection of Molecular Biotargets
Vanessa Cupil-Garciaab, Pietro Strobbiaac, Yang Ranad, Bridget M. Crawfordac, Hsin-Neng Wangac, Rodolfo Zentelle, Tai-Ping Sune, and Tuan Vo-Dinhabc
a. Fitzpatrick Institute for Photonics, Duke University, Durham, NC 27708, USA,
b. Department of Chemistry, Duke University, Durham, NC 27708, USA,
c. Department of Biomedical Engineering, Duke University, Durham, NC 27708, USA,
d. Guangdong Provincial Key Laboratory of Optical Fiber Sensing and Communications, Institute of Photonics Technology, Jinan University, Guangzhou 510632, China,
e. Department of Biology, Duke University, Durham, NC 27708, USA.
View abstract
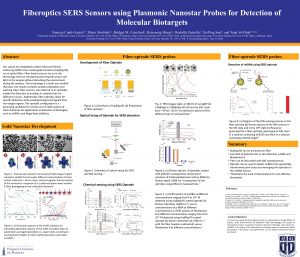
Our group has integrated surface-enhanced Raman scattering (SERS) silver coated gold nanostars on an optical fiber. Fiber-based sensors are an in situ technology that can simultaneously bring the sensor and light to the sample without disturbing the environment during the analysis. This technology is a multi-use method that does not require complex sample preparation and washing steps. Fiber sensors, also referred to as optrodes, enable the detection of analytes in samples that are difficult to access. Additionally, fiber-optrodes allow for specific detection while evading background signals from non-target regions. The fiber-optrode was used to detect miRNA and illegal food additives, such as Rhodamine B. Our group functionalized the tip of the fiber-optic with inverse molecular sentinel nanosensors. The fiber-optic biosensor was used to detect miR156 from leaf tissue of N. benthamiana. It is desirable to develop methods for field detection of miR156 since it is involved in the flowering of plants which is important in regulating plant growth and improving the production of biomass. The fiber-optic sensor was capable of only binding the target miRNA even in the presence of other miRNA. This optrode configuration is a promising candidate for remote and in-field analysis of trace chemicals and biotargets.
28. A Generative Adversarial Network for Artifact Removal in Photoacoustic Computer Tomography with a Linear-array Transducer
View abstract
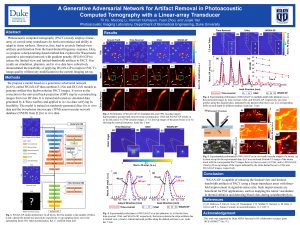
Photoacoustic imaging (PAI), especially photoacoustic computed tomography (PACT), is quickly growing in clinical application. For many of its implementations, non-full-view (linear-array, curved-array, etc.) transducers are used for both convenience and ability to adapt to tissue surfaces. However, these probes have significant drawbacks. They lead to severely limited-view artifacts (LVA) and distortion from the band-limited frequency-response (BLFR) of the transducer. Existing works in both hardware and software domain are not sufficient for solving these problems, especially for in vivo data. In this paper, we propose a deep-learning-based method named PI-Net that utilizes recent advances in generative adversarial networks to address LVA and BLFR artifacts, while improving resolution by four times. Preliminary results in simulated, phantom, and in vivo trials represent the potential of this model’s application to in vivo PACT reconstruction. Qualitatively and quantitatively, PI-Net significantly corrects LVA and BLFA, while increasing the resolution by four times and the contrast-to-noise ratio by six times for in vivo data.
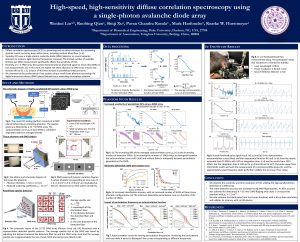
29. High-speed, high-sensitivity diffuse correlation spectroscopy using a singlephoton avalanche diode array
View abstract
Diffuse correlation spectroscopy (DCS) measures rapid changes in scattered coherent light to detect blood flow and functional dynamics within tissue. While its sensitivity to minute scatterer displacements leads to a number of unique advantages as compared to other technologies, conventional DCS systems become photon-limited when attempting to probe deep into tissue, which leads to long measurement windows (1 sec) and low temporal resolution. Here, we present a high-sensitivity DCS system that uses 1024 parallel detection channels provided by a 32×32 single-photon avalanche diode (SPAD) array. We demonstrate that this new parallelized measurement strategy is sensitive to mm-scale perturbations up to 1 cm within a tissue-like phantom at up to 100 Hz sampling rate, and also show that it can measure high-fidelity human pulse signals from the forehead with a temporal resolution of 33~ms. These results suggest that a highly parallelized approach to DCS can overcome standard sensitivity and temporal resolution limits to open up new applications for measuring high-speed biological signals.
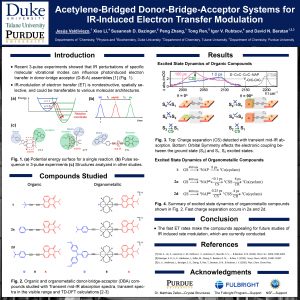
30. Acetylene-bridged Donor-Bridge-Acceptor Systems for IR-Induced Electron Transfer Modulation
Jesús Valdiviezo1, Xiao Li4, Susannah D. Bazinger5, Peng Zhang1, Igor V. Rubtsov4, Tong Ren5, David N. Beratan1,2,3
1. Department of Chemistry, Duke University, Durham, NC, United States.
2. Department of Physics, Duke University, Durham, NC, United States.
3. Department of Biochemistry, Duke University, Durham, NC, United States.
4. Department of Chemistry, Tulane University, New Orleans, LA, United States.
5. Department of Chemistry, Purdue University, West Lafayette, IN, United States.
View abstract
The efficiency of opto-electronic devices strongly depends on the photoinduced charge separation and charge recombination rates. Current strategies to control electron transfer kinetics have reported to be chemically invasive or irreversible. In recent years, UV-IR-Vis-3-pulse experiments have indicated that IR perturbations of a specific molecular vibrational mode can influence photoinduced electron transfer rates in donor-bridge acceptor (D-B-A) assemblies. The possibility to use infrared excitations to control electron transfer is of substantial interest, since it is a nondestructive method (the energy absorbed ranges between 2.5-8.5 kcal/mol), spatially selective (specific bonds can be excited) and infrared active functional groups can be easily incorporated into molecular architectures. Through a collaborative effort between experiment and theory, we combined synthesis, computational methods and ultrafast spectroscopy techniques to investigate the evolution of excited states in organic and organometallic D-B-A complexes. Under the presence of strong donors, the optical excitation of the acceptor leads to ultrafast formation of charge separated states (CSS) These fast dynamics makes them excellent candidates for electron transfer rate modulation by infra-red excitation. Moreover, it is found that the excitation of the CºC stretching mode of the bridge slows the rate of CSS formation, providing evidence of the possibility to use IR pulses to modulate the electron transfer rate in compounds without rare-earth metals.
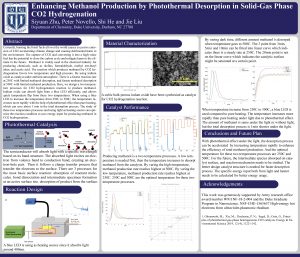
31. Enhancing Methanol Production by Photothermal Desorption in Solid-Gas Phase CO2 Hydrogenation
View abstract
Currently, burning the fossil fuels all over the world causes excessive emission of CO2 accelerating climate change and causing detrimental harm to the environment. The capture of CO2 and converting it into a high-value fuel has the potential to close the carbon cycle and mitigate harm to the climate in the future. Methanol is widely used in the chemical industry for producing chemicals, such as olefins, formaldehyde, methyl tert-butyl ether, and acetic acid. The reaction which produces methanol by CO2 hydrogenation favors low temperature and high pressure. By using indium oxide as catalyst under ambient atmosphere. There is a fastest reaction rate at 250C with limited methanol desorption, and fastest methanol desorption at 500C with limited methanol production. Here, we design a two temperature processes for CO2 hydrogenation reaction to produce methanol. Indium oxide can absorb light from a blue LED efficiently, and allows quick temperature flip from these two temperatures. When using a blue LED to increase the temperature from 250C to 500C, the temperature increases more rapidly with the help of photothermal effect than pure heating, which can save about 1 min in the total desorption process. The study of these two temperature processes and using light as heating source can optimize the reaction condition to save energy input for producing methanol in CO2 hydrogenation.
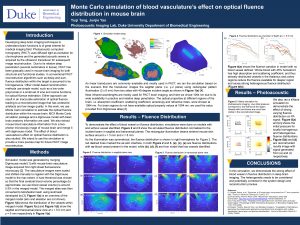
32. Monte Carlo simulation of blood vasculature’s effect on optical fluence distribution in mouse brain
View abstract
Developing deep brain imaging techniques to understand brain functions is of great interest for medical imaging field. Photoacoustic computed tomography (PACT) uses diffused light as excitation for chromophores and the generated acoustic waves is acquired by the ultrasonic transducer for subsequent image reconstruction. Due to its relative deep penetration (several centimeters in tissue), PACT has been gradually used in mouse brain imaging for both structural and functional studies. In conventional PACT reconstruction algorithms such as delay-and-sum, fluence distribution within the target is assumed to be homogeneous. Other model-based reconstruction methods use simple model, such as a low order polynomial or a small set of sine and cosine functions, for optical fluence estimation. Either approach can result in an inaccurate estimation of optical fluence, leading to a reconstructed image that has undesired artefacts and low image quality. In this work, we use Monte Carlo simulation to estimate the optical fluence distribution within the mouse brain. MCX Monte Carlo simulation package and a digimouse model with basic brain anatomy information are used. We also extract mouse blood vasculature information from a two-photon microscopy image of mouse brain and merge it with digimouse model. The effect of blood vasculature’s effect on optical fluence distribution is also studied. We intend to use this simulation to provide a more precise map for future PACT image reconstruction.
33. Improving SNR of Photoacoustic Imaging Using a Matched Filter
View abstract
Matched filtering is widely used in digital communications, radar and sonar detection, as well as gravitational-wave astronomy, in order to improve SNR substantially. In our method, we cross-correlate the template signal of the system’s impulse response with the raw photoacoustic signals, which can suppress the uncorrelated detection noise and signal interference and thus improve the SNR of the PA images. We obtained the impulse response of our photoacoustic imaging system and validated our method on the mouse ear vascular image. Our results have demonstrated SNR improvements in PA imaging by rejecting strong background noises.
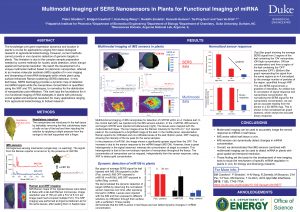
34. Multimodal Imaging of SERS Nanosensors in Plants for Functional Imaging of miRNA
Pietro Strobbia1,2, Bridget M. Crawford1,2, Hsin-Neng Wang1,2, Rodolfo Zentella3, Maxim I. Boyanoa3, Tai-Ping Sun4, Kenneth M. Kemner3, Tuan Vo-Dinh1,2,5
1 Fitzpatrick Institute for Photonics, Duke University
2 Department of Biomedical Engineering, Duke University
3 Department of Biology, Duke University
4 Department of Chemistry, Duke University
5 Biosciences Division, Argonne National Laboratory
View abstract
Monitoring gene expression within whole plants is critical for many applications ranging from plant biology to agricultural biotechnology 13 and biofuel development; however, no method currently exists for in vivo monitoring of genomic targets in plant systems. We developed a unique multimodal method based on plasmonic nanoprobes capable of in vivo imaging and biosensing of microRNA biotargets within whole plant leaves by integrating three different and complementary techniques: surface-enhanced Raman scattering (SERS), X-ray fluorescence (XRF), and plasmonics-enhanced two-photon luminescence (TPL). The method developed uses plasmonic nanostars, which not only provide large Raman signal enhancement, but also allow for localization and quantification by XRF and plasmonics-enhanced TPL, owing to gold content and high two-photon luminescence cross-sections. Our method uses inverse molecular sentinel (iMS) nanoprobes for SERS bioimaging of microRNA within Arabidopsis thaliana leaves to provide a dynamic SERS map of detected microRNA targets while also quantifying nanoprobe concentrations using XRF and TPL. The nanoprobes were observed to occupy the intercellular spaces upon infiltration into the leaf tissues. This report lays the foundation for the use of plasmonic nanoprobes for in vivo functional imaging of nucleic acid biotargets in whole plants, a tool that will revolutionize bioengineering research by allowing the study of these biotargets with previously unmet spatial and temporal resolution.
© 2026 Frontiers in Photonics Science & Technology
Theme by Anders Noren — Up ↑